This week we review a disease called Alzheimer-related primary progressive aphasia (PPA-AD). It is well known that a primary symptom of Alzheimer’s disease (AD) is memory impairment, while the primary symptom of primary progressive aphasia (PPA) is an isolated language disturbance. Two thirds of PPA cases are caused by a tauopathy called Frontotempral degeneration (FTD) but the remaining third are due to AD pathology making up the PPA-AD group.
The fact that PPA-AD has only language dysfunction as opposed to memory impairment has perplexed researchers for years. Recently, a group found a possible explanation for this during a longitudinal follow-up on a cohort (n = 31) with AD, divided into typical amnestic presentations (n = 14) versus language presentation (PPA-AD, n = 17). Participants received longitudinal memory and language tests, biomarker analyses, and the majority agreed to autopsy. Using these metrics, they discovered some trends that may shed light on why PPA-AD and typical AD dementia (DAT-AD) have such different clinical presentations.
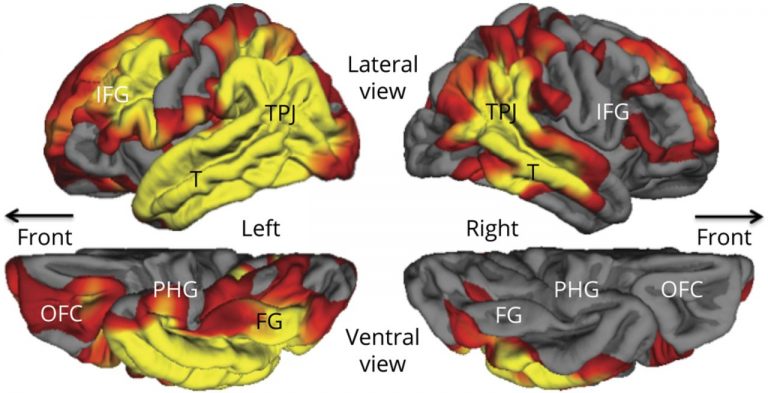
In the PPA-AD group, participants had a significant yearly decline of 6.21% in object naming scores and 4.25% for a measure of global language performance, with no significant decrease in memory. Those in the DAT-AD group, meanwhile, had a significant yearly decrease of 2.15% for memory and 4.05% for object naming. Structural imaging was also done on the PPA-AD group at their initial visits showing cortical thinning, especially of the language-dominant left hemisphere, extending throughout the language network. Significant thinning of the parahippocampal gyrus was present only on the left side (as shown below, denoted by PHG).
To assess neuropathology, Aβ plaques, neurofibrillary tangles (NFTs), and overall plaque density were quantified. The PPA-AD group showed maximum levels of Aβ and NFTs, though researchers focused on NFTs as their distribution and density more strongly correlates to cognition. Specifically, the PPA-AD group had severe NFT pathology in the neocortex and all medial-temporal lobe structures associated with memory. However, 2 of the 8 who agreed to undergo autopsy were found to be of the “hippocampal-sparing type” where cortical NFT density is higher than in memory-related structures. Interestingly, despite the decrease in hippocampal NFT density and sparing of memory, those 2 participants had severe NFT-induced degeneration of memory regions. Furthermore, bilateral comparisons revealed that the left hemisphere of these 2 participants had more Aβ plaques while other patients had elevated NFTs, suggesting a dichotomy between pathogenesis of hippocampal-sparing type PPA-AD and typical PPA-AD.
This leaves the question, what induces resilience of memory structures in PPA-AD? Well, in PPA-AD the hippocampal gyrus primarily degenerates on the left hemisphere. Previous lesion studies on this area have shown that episodic memory function only significantly declines when bilateral lesioning or degeneration of hippocampal structures occurs. However, while this study showed decreased NFT aggregation in memory-related structures in PPA-AD compared to cortical areas, previous studies have had varied results indicating that further longitudinal studies of this type are required.
One other possible explanation for memory resilience in PPA-AD involves APOE status. The PPA-AD group had a 14.7% incidence of ε4 alleles, which inhibits neuronal plasticity, while the rest had ε3 alleles, which enhance neuronal plasticity. Interestingly, this frequency of ε4 alleles matches control populations with PPA-AD, while the DAT-AD group had a 42% ε4 frequency with 3 homozygous carriers, almost twice as high as control populations. Thus ε4 presence likely increased vulnerability of memory networks in the DAT-AD group by inhibiting compensatory mechanisms like neuroplasticity, while those with PPA-AD, having reduced ε4 and increased ε3 frequency, may increase resilience to neurodegeneration by allowing for neuronal plasticity.